Phage display peptide libraries have emerged as powerful tools in molecular biology, drug discovery, and therapeutic development. These libraries offer an efficient means of identifying peptides that bind specifically to a variety of targets, such as proteins, antibodies, or other biomolecules. The ability to rapidly generate and screen vast libraries of peptides has revolutionized our approach to drug development, biomarker discovery, and the study of protein interactions. In this article, we will explore the methods of constructing phage display peptide libraries and their broad applications in scientific research and medicine.
What is Phage Display?
Phage display is a technique that allows the display of peptides, proteins, or antibodies on the surface of bacteriophages (viruses that infect bacteria). These phages serve as a vehicle for the presentation of peptides, which are encoded by the genetic material of the phage. The technique allows researchers to link the genetic information of a peptide to its physical display on the phage surface, making it possible to screen large libraries of peptides for interactions with specific targets.
In a typical phage display experiment, a peptide or protein of interest is fused to a coat protein of the phage, making the peptide “displayed” on the outer surface. When the phage is exposed to a target molecule, those phages that bind to the target are isolated and amplified, enabling researchers to identify specific peptides with high affinity for the target.
Methods of Phage Display Peptide Library Construction
Phage display peptide library construction involves several key steps that enable the generation of diverse peptide libraries, which can then be screened for binding to specific targets. Below are the most common methods used to construct these libraries.
1. Library Design and Synthesis
The first step in phage display library construction is designing the peptide library. The diversity of the library is determined by the number and type of amino acids included in the peptide sequences. Libraries are typically designed to represent random peptide sequences, which may vary in length and composition depending on the specific research goals.
- Random Peptide Libraries: In random peptide libraries, peptide sequences are generated with a randomized codon set, allowing for the generation of a large variety of peptides. These libraries can contain millions to billions of different peptide sequences.
- Predefined Peptide Libraries: In some cases, libraries are designed with predefined peptide motifs or sequences that are believed to be of particular interest, such as epitopes from antigens, or motifs from known binding proteins.
2. Phage Display Vector Construction
The next step involves inserting the synthesized peptide sequences into a phage display vector. A phage display vector typically consists of a phage genome that can be genetically engineered to express the peptide of interest on its surface. The most commonly used bacteriophages for display are M13 phage and T7 phage, though other types of phages can also be used.
- Fusion Proteins: The peptide is usually fused to a coat protein of the bacteriophage, such as the gene III protein in M13 phages. This fusion allows the peptide to be expressed on the surface of the phage, making it accessible for interaction with potential binding targets.
- Vector Construction: The peptide gene is inserted into the phage genome using molecular cloning techniques, such as restriction enzyme digestion and ligation, or more modern approaches like gateway cloning or Gibson assembly.
3. Phage Display Library Packaging
Once the phage display vector has been constructed, the next step is packaging the recombinant phage. This involves infecting bacteria (typically Escherichia coli) with the phage vectors, which allows the production of phages containing the newly inserted peptide sequences. The bacteria then replicate and produce large numbers of phages, each displaying a unique peptide on its surface.
- Infection of E. coli: The recombinant phage vectors are introduced into a host bacterium, such as E. coli, which will then produce and display the phage-peptide fusion proteins.
- Amplification: The infected bacteria are cultured, and phages are harvested from the bacterial culture supernatant, where they can be purified for subsequent use in screening assays.
4. Library Enrichment and Screening
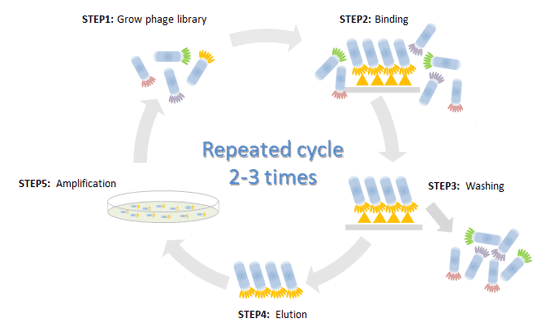
After the library is prepared, the next step is to screen for peptides that bind to the desired target. This is typically done using a process known as biopanning, where the phage library is incubated with the target (such as a protein, antibody, or cell surface). The phages that bind to the target are then isolated, amplified, and subjected to further rounds of selection, known as panning, to enrich for high-affinity binders.
- Selection for Specific Binding: After each round of panning, the unbound phages are washed away, and the bound phages are eluted and amplified. This process is repeated for several rounds to increase the affinity and specificity of the peptides for the target.
- Elution and Amplification: The peptides that remain bound to the target are eluted, typically by changing the conditions (e.g., using high salt or acid), and the phages are amplified by infecting fresh E. coli.
5. Sequencing and Analysis
Once the final round of panning has been completed, individual phage clones are picked, and the sequences of the displayed peptides are determined through DNA sequencing. The peptides that consistently bind to the target are identified, and their sequences can be further analyzed for functional or structural motifs that are relevant for the biological process under study.
- Sequence Analysis: The obtained peptide sequences can be analyzed to identify common motifs or specific amino acid residues critical for binding, providing valuable insights into the mechanism of interaction with the target.
- Further Testing: The identified peptides may be synthesized and tested for their biological activity, including binding affinity, specificity, and potential therapeutic effects.
Applications of Phage Display Peptide Libraries
Phage display peptide libraries have a wide range of applications in basic research, drug development, and biotechnology. Some of the most significant applications include:
1. Antibody Development
Phage display is commonly used for the development of therapeutic antibodies. By screening peptide libraries against a target protein, researchers can identify peptide sequences that mimic antibody epitopes. These peptides can then be used as starting points for the design of monoclonal antibodies or other protein-based therapeutics.
- B-cell Epitope Mapping: Phage display can help identify B-cell epitopes that antibodies recognize, assisting in vaccine design and diagnostics.
2. Targeted Drug Discovery
Phage display peptide libraries are often used to discover peptides that bind specifically to disease-related targets. These peptides can serve as leads for the development of peptide-based drugs or can be used as scaffolds for designing small molecule inhibitors.
- Inhibitor Development: Identified peptides can be optimized to create potent inhibitors of enzymes or other proteins involved in disease processes.
3. Biomarker Identification
Phage display can be employed to discover biomarkers for diseases such as cancer, autoimmune disorders, and infectious diseases. Peptides that bind specifically to disease-associated biomarkers can be used for diagnostic purposes or therapeutic interventions.
- Cancer Diagnostics: Phage display libraries are often used to identify peptides that bind to tumor-associated markers, providing a basis for non-invasive cancer detection.
4. Vaccine Development
Phage display peptide libraries are also used to identify epitopes for vaccine development. By displaying peptides that mimic viral or bacterial proteins, researchers can stimulate immune responses that protect against infection.
- Epitope-Based Vaccines: Phage display enables the identification of potential vaccine candidates by selecting peptides that elicit strong immune responses.
5. Protein-Protein Interaction Studies
Phage display can be used to study protein-protein interactions, which are crucial for understanding cellular signaling, gene regulation, and disease mechanisms. Peptides that bind to specific protein interfaces can provide insights into how proteins function and interact in vivo.
- Drug Target Discovery: Peptides from phage display libraries can help discover novel protein-protein interactions that can be targeted for drug development.
Conclusion
Phage display peptide library construction has revolutionized molecular biology and biotechnology, offering a powerful tool for discovering new peptides, proteins, and therapeutic agents. Through the process of library construction, phage display enables the screening of vast numbers of peptides for specific binding to targets, accelerating drug discovery, biomarker identification, and vaccine development. As the field continues to evolve, advances in phage display technologies promise to unlock even greater possibilities in medical and scientific research.
Table of Contents